- Published on
Benchmark of popular graph/network packages
- Authors
- Name
This post is superseded by an updated benchmark
In this post I benchmark the performance of 5 popular graph/network packages. This was inspired by two questions I had:
Recently, I have been working with large networks (millions of vertices and edges) and often wonder what is the best currently available package/tool that would scale well and handle large scale network analysis tasks. Having tried out a few (networkx in Python and igraph in R) but on different problems, I thought it would be nice to have a head to head comparison.
Running large scale computations is also much easier nowadays with the availability of virtual machines that could be easily spinned up thanks to the growth of cloud computing. I think the trend of powerful single machines will eliminate a lot of the need for enterprise clusters so it will be interesting to see how far we can push a single machine using optimised algorithms.1
To replicate the benchmark study and for the full codes, please refer to my github repository. Instructions on how to setup and install the packages are also located in the repository.
Setup
The benchmark was carried out using a Google Compute n1-standard-16 instance (16vCPU Haswell 2.3GHz, 60 GB memory). I compare 5 different packages:
Networkx is written in Python while the other four packages are based on C / C++ but have Python APIs. Igraph has a R and Mathematica binding as well but to be consistent the following benchmark was based on the Python one. The other 3 libraries (snap, networkit and graph-tool) have an additional emphasis on performance with multi-processing capabilities built in.
Selecting what tasks to compare on is not really a trivial task with each package offering various tools and capabilities. In the end, I decided to focus on 5 specific problems:
- loading the data
- single source shortest path
- page rank
- k-core decomposition
- strongly connected components
Loading is more of an I/O task while the other 4 are common graph algorithms. Disclaimer: I try as much as possible to specify the same parameters for each algorithm but differences in API across the packages could translate to actual differences in how the algorithm is run and the final output.
13/12/2019 Edit: Some of the observed differences in performance might be a result of different stopping criteria used - see algorithms for more information.
3 datasets from the Stanford Large Network Dataset Collection were used in the exercise:
While it is the easiest to rank the packages based on the run-time of the algorithms, it is only one of the many considerations of what makes a good package. I try to offer a more subjective view based on my experience with these packages.
Related Benchmarks
Here's a list of other comparative benchmarks for the interested viewer to check out:
- Graph-tool performance comparison, compares graph-tool with igraph and networkx
- SNAP research paper, compares snap with igraph and networkx
- Networkit technical paper, compares networkit with igraph and graph-tool
Most of them were written in 2015/2016 and it will be interesting to see if anything has changed since then.
Results
All timings reported are normalised to reflect the run time for a single run of the task.
Networkx is much slower than any of the other libraries. Across all computation tasks and for all datasets it is around 10 times slower than the slowest library.2 For example, it took 67s to run the single source shortest path problem on the Pokec dataset compared to 6.8s for networkit (the next slowest). Page rank took more than 10 minutes to run compared to 1 minute for igraph. Hence, I left it out of the comparison plots.
Here are the run times of the remaining four packages:
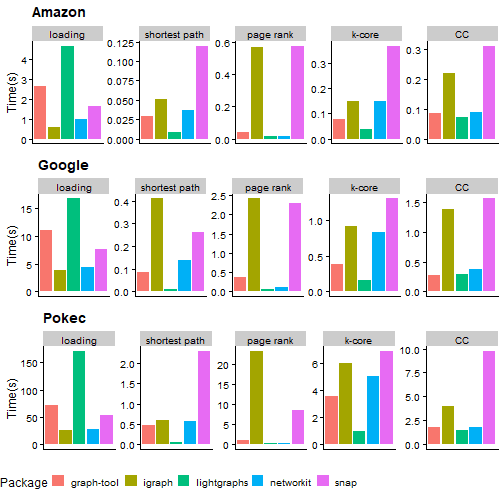
Full results can be seen from the table below:
| dataset | Algorithm | graph-tool | igraph | networkit | networkx | snap |
|---|---|---|---|---|---|---|
| Amazon | connected components | 0.09 | 0.48 | 0.21 | 5.94 | 0.40 |
| Amazon | k-core | 0.11 | 0.33 | 0.01 | 8.62 | 0.42 |
| Amazon | loading | 5.00 | 0.79 | 3.27 | 9.96 | 1.90 |
| Amazon | page rank | 0.05 | 1.59 | 0.01 | 25.71 | 0.90 |
| Amazon | shortest path | 0.06 | 0.12 | 0.32 | 3.31 | 0.14 |
| connected components | 0.32 | 2.23 | 0.65 | 21.71 | 2.02 | |
| k-core | 0.57 | 1.68 | 0.06 | 153.21 | 1.57 | |
| loading | 67.27 | 5.51 | 17.94 | 39.69 | 9.03 | |
| page rank | 0.76 | 5.24 | 0.12 | 106.49 | 4.16 | |
| shortest path | 0.20 | 0.69 | 0.98 | 12.33 | 0.30 | |
| Pokec | connected components | 1.35 | 17.75 | 4.69 | 108.07 | 15.28 |
| Pokec | k-core | 5.73 | 10.87 | 0.34 | 649.81 | 8.87 |
| Pokec | loading | 119.57 | 34.53 | 157.61 | 237.72 | 59.75 |
| Pokec | page rank | 1.74 | 59.55 | 0.20 | 611.24 | 19.52 |
| Pokec | shortest path | 0.86 | 0.87 | 6.87 | 67.15 | 3.09 |
I/O
Looking at the plots above, graph-tool and networkit loads data much more slowly than the other two libraries. I was reading the datasets as a tab delimited file and graph-tool basically uses a Python code to parse the input. The other 3 packages should be using C libraries to read the files which result in better performance.3
Algorithms
Networkit and graph-tool takes the top spot in most of the tests with graph-tool having the shortest run time for the single source shortest path and connected components problems and networkit winning the race for k-core and page rank.
When networkit is fast, it is extremely fast. On the pokec dataset it takes just 0.2s to run the page rank algorithm (graph-tool: 1.7s, igraph: 59.6s, snap: 19.5s). For the k-core decomposition it is also 10 times faster than all other competitors or 2000 times networkx. That is consistent with the findings of their research paper where they claim that using some of the latest state of the art algorithms led to their processing speed being faster by an order of magnitude. However, for the shortest path problem (not analysed in their paper) it lags behind all other packages.4
13/12/2019 Edit: Matthew Galati from SAS pointed out that for the pagerank algorithm, networkit (as of version 6.0) uses L2 norm as a stopping criteria while other packages use the L1 norm. This means that it is doing fewer iterations and the speed is somewhat artificial. Thanks Matthew!
graph-tool is the most steady performer and achieves very impressive performance across all four tasks. With openMP support it betters igraph and snap across all tasks. It is 3 to 10+ times faster than those two packages.
igraph and snap achieves mostly similar performance across all tasks with a slight edge towards snap. This is also consistent with snap's research findings.
Other Considerations
There are also other important considerations when making a switch from networkx or igraph to one graph-tool or networkit.
Packages
First, the algorithms available differ quite significantly across the packages. Users interested in switching to one of these packages should read the documentation on the list of features available. For example, while they all contain the basic tools needed to manipulate networks, graph-tool does not have the more usual modular clustering tools but has additional functionalities on statistical inference on graphs using stochastic block models.
Visualising networks is also an important part of the analytical tool chain. Igraph implements quite a few layout algorithms and renders them using the cairo library. Snap supports graphviz while graph-tool supports both graphviz and cairo. Networkit takes a different approach and relies on networkx to draw while also providing support and integration with Gephi via its streaming plugin.
API
Moving away from native Python or R means that the syntax for the packages can sometimes be quite convoluted. I compare the syntax for the shortest path problem below. Writing it in networkx would look something like this:
nx.shortest_path_length(g, nodeid)
igraph:
g.shortest_paths([g.vs[node_index]])
graph-tool:
shortest_distance(g, g.vertex(node_index))
networkit:
distance.BFS(g, node_index).run()
snap:
NIdToDistH = snap.TIntH()
snap.GetShortPath(g, node_index, NIdToDistH, True)
Of all, I find snap's the most cumbersome since one has to define an additional variable (with the correct variable type) to store the results before running it. Running more advanced functions on graph-tool and networkit also requires a user to pre-define variables with the correct type to store results.5
Support and Documentation
User support and documentation is really important when one wants to use the project in an actual project setting. Networkx is by far the winner in this category with more than 4k github stars and lots of issues documented in github and stackoverflow. Igraph fairs decently as well with more than a thousand stars across its different modules.
graph-tool and networkit has much smaller followings though the creators seem relatively responsive to user issues and the packages are in active development.
snap was last updated on July 2018 but still supports only Python 2.7.x versions.
Conclusion
Overall, I am pleasantly surprised at the performance of the libraries especially graph-tool and networkit and plan to play around with them further. The fact that they breeze through the Pokec dataset is a good sign, but it will be interesting to find out what is the limit before computation becomes slow or memory issues start appearing.
As for recommendations on which package people should learn, I think picking up networkx is still important as it makes network science very accessible with a wide range of tools and capabilities. If analysis starts being too slow (and maybe that's why you are here) then I will suggest taking a look at graph-tool or networkit to see if they contain the necessary algorithms for your needs.
Footnotes
A 240 GB machine with 64 vCPUs currently cost about $1.5k USD / month on Google Cloud Compute. While relatively expensive, one can spin up the machine just for very heavy computation tasks and shut it down once the task is complete. Such intensive computation once reserved for enterprises and research institutions can now be replicated by almost anyone. ↩
After trying it out for 1 test run, I run the profiling tests only 10 times fewer when using the networkx library. ↩
Graph-tool should read from other file types such as graphml or dot much faster though I did not actually try it out. ↩
I used the BFS search to proxy for the single source shortest path problem as there does not seem to be a direct API call for it. ↩
I guess this is required of statically typed languages like C and C++. ↩